Author(s): Mbugua J K*, Mbui D N, Mwaniki J M and Waswa a G
Biogas can be produced from vegetable and fruit wastes from marketplaces, inoculated with slaughterhouse waste at different temperature ranges. In the current study, biochemical methane potential of twenty different market wastes were calculated using a web-based application developed using R programming language. The proximate analysis of the substrates was carried out using standard procedures, while the microbial community in the rumen fluid used as inoculum was done using standard plate method. The substrates digestibility was calculated using COD, volatile matter and lignin contents.The theoretical biogas generated from market wastes was obtained using macro-molecular composition i.e., crude fat, proteins, fiber and carbohydrates loaded to online biogas application.
The proximate composition of the substrates showed that carbohydrates levels was higher in all the feed-stocks compared to proteins and fats. The moisture levels were in the range of 74.31-95.86% for all the wastes. Low percentages of proteins and fats were observed at 0.52 -3.49% and 0.09-1.54%, respectively. The theoretical methane obtained from the market wastes was higher in avocado at 65.52 % and lowest in cabbage at 50.90%.
The results obtained indicates that the methane potential of market waste heavily depend on the macro-nutrient composition of the substrate. The study recommends application of online based application to investigate the methane potential of substrates before carrying out the experiment in the laboratory.
Anaerobic degradation of organic matter to renewable energy entails lab and pilot scale experiments, theoretical calculations, process simulations and modeling [1]. has developed a biogas software tools which measures and predict methane production for a given substrate. Laboratory scale involves biochemical methane potential studies to predict maximum methane production from a substrate. Relationship between BMP and experimental data employs modeling and simulation calculations [2]. Proximate composition of vegetable and fruits wastes can be used to predict methane production [3]. The methane potential of market wastes can also be predicted by analyzing the substrate contents before loading to the digester as reported by [4].
Biogas estimation applications are software tools or models that are used to estimate the potential biogas production from different organic waste materials [5]. These applications are designed to predict the amount of methane gas that can be produced from various substrates such as agricultural waste, animal manure, food waste, and municipal solid waste. Some common biogas estimation applications include:
Anaerobic Digestion Model (ADM): ADM is a widely-used biogas estimation application that is used to simulate the anaerobic digestion process. ADM uses mathematical equations to predict the amount of biogas that can be produced from different types of organic waste materials [6-8].
BioWin: BioWin is a software application that is used to model and simulate wastewater treatment processes. It can be used to estimate biogas production from organic waste materials that are generated during the wastewater treatment process [9-11].
Biogas ax: Biogas ax is an online biogas estimation tool that is designed to estimate the biogas production potential from different types of organic waste materials. It uses a database of empirical data to estimate the amount of biogas that can be produced from specific waste types [12].
Bio Gasol: Bio Gasol is a biogas estimation tool that uses a thermodynamic model to estimate the potential biogas production from different types of organic waste materials. It can be used to estimate biogas production from various substrates such as agricultural residues, food waste, and sewage sludge [13].
AD-Fit: AD-Fit is a biogas estimation tool that uses statistical models to predict the biogas production potential from different types of organic waste materials. It can be used to estimate biogas production from various substrates such as animal manure, food waste, and energy crops [14].
BioTool: an easily manageable tool for end-users to predict the biogas production rate to ease the transition to a flexible and proactive operation of digesters at both biogas plants as well as at wastewater treatment plants. To provide a high flexibility to end-users but keep BioTOOL easily manageable, the approach uses a pre-trained artificial neural networks library consisting of different variable combinations. BioTOOL uses a seek-andimplement method to find the correct stored network for the entered input. This approach results in predictions similar to the optimized per-trained network. It has been demonstrated that the idea of a flexible prediction tool could be fully realized [15].
These are just a few examples of the many biogas estimation applications that are available. Each application has its own set of features and capabilities, and the choice of application will depend on the specific needs of the user [16]. For example, a spreadsheet was created to estimate biogas production and other operational factors using substrate loading rates and temperature settings. This user-friendly program could be used as a decision-support tool to provide recommendations for some working conditions and estimation of the biogas yield under different scenarios [17].
carried out a study aimed at modeling bio-digestion systems as a function of the most influencing parameters to generate two robust algorithms on the basis of the machine learning algorithms, including adaptive network-based fuzzy inference system and least square support vector machine [18]. The models are assessed utilizing multiple statistical analyses for the actual values and model outcomes. Results from the suggested models indicate their great capability of predicting biogas production from vegetable food, fruits, and wastes for a variety of ranges of input parameters [18].
Described a model made in R programming environment used for prediction of methane production potential using biogas package [1,19,20]. Detailed description of biogas package use in predicting methane and biogas production can be obtained from https:// cran.r-project.org/package=biogas.
Online biogas application (OBA) is an online server-based application developed to process and predict biogas prediction. Calculations are run using biogas package in R programming language. The package was developed by Sasha D. Hafner and Charlotte Rennuit while some work was done by Jon Katz.
Methane production can be predicted based on three main substrates characteristics. These are chemical oxygen demand, empirical (chemical) formula or macromolecular composition [21]. The application output methane standardized at 1atm and the percentage methane. In this study, macromolecular substrate composition is used to predict biogas production from twenty different market wastes.
The fruits and vegetable waste used as substrates were obtained from Kangemi and Wakulima market in Kiambu County while rumen fluid used as inoculum was obtained from Dagoretti slaughterhouse. The sampling sites are shown in figure 1.
Figure 1: A map of sampling sites
Fresh solid vegetable and fruits market wastes; Cabbage (Brassica oleracea capittta), Coriander (Coriandrum sativum.), Spinach (Spinacia oleracea), Kales (Brassica oleracea acephala), Pumpkin Leaves (Cucurbita maxima), Kahurura (Cucumis ficifolia), Pig Weed (Amaranthus spp.), African Nightshade (Solanum nigrum), Togotia (Erucastrum arabicum), comfrey (Symphytun officinale), Banana (Musa spp), Sweet Potato (Ipomoea batatas), Cucumber(Cucumis sativus), Watermelon (Citrullus lanatus), Tomato (Lycopersicon lycopersicum), Potato (Solanum tuberosum), Avocado (Persea americana), Mango (Mangifera indica), Papaya (Carica papaya), and Courgette (Cucurbita pepo) were sliced into small pieces with a knife and then blended for toxic substances and macro and micro nutrient and heavy metals analysis and proximate analysis studies.
Ash, moisture and fiber contents were determined using method [22]. Fat, crude nitrogen and protein contents were determined using Soxhlet extraction and micro-Kjedhal methods described in method [23]. Carbon content was carried out using method, Energy content was carried out using the AOAC method described by while Total and Volatile solids were determined using method [24-26].
The sample was heated at 600°C for 1 hour in a muffle furnace, cooled, and afterwards weighed to determine the amount of ash present. For 2-4 hours, one gram of each sample was burned at 550°C. The ash levels were calculated using equations 1.0 and 2.0.

The weights of the crucible and ash, blank crucible, and sample before burning are represented as W3, W1, and Ws, respectively
The Kjeldahl technique was used to assess the protein in the samples [27]. In order to digest roughly 0.5-1.0 g of dry waste samples, Sulphuric acid was heated along with a digestion mixture that included potassium sulphate and the selenium catalyst. To make the digested mixture alkaline, 0.1M sodium hydroxide was added. The end-product was ammonium sulphate. Before titrating against standard hydrochloric acid, ammonia was collected in a solution of 2% boric acid. The equations 3.0 and 4.0 were used to calculate the total protein.

Where V is the volume used for distillation, S is the sample titration value, B is the blank titration reading, N is the HCl’s normality, D is the sample’s dilution upon digestion, and N is the milli equivalent weight of nitrogen.
Through using the Soxhlet apparatus, the ether extract method was used to determine the total crude fat in the samples. Approximately 1.5 to 2.5 g of the dried samples were placed in a fat-free thimble, covered in filter paper, and thereafter put into the extraction tube. The material was weighed into a receiving beaker that had been cleaned, dried, and filled with petroleum ether before the extraction apparatus was put together. The extraction procedure got underway. The ether was expelled after 4-6 siphoning, and the beaker was then detached before the final siphoning. The ether was then vaporized while the extract was moved to a clean glass dish in a water bath. The dish was desiccated after drying at 105°C for two hours. The total crude fat was then calculated using equation 5.0. Ws is the combined weight of the crucible as well as the sample.
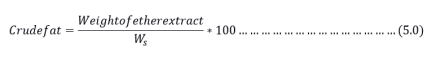
The digestibility methods that had been indicated earlier, assumed that all the degradable matter is broken-down; as a consequence, a compensation of this assumption is made by the experimental BMP examination. The elemental biodegradability (BD ele ) of the substrate was estimated by equation 6.0 as described by [28].

The theoretical biochemical methane potential of 20 market wastes was investigated using biogas online application by [1]. The application which is built in R programming language is found at https://cran.r-project.org/package=biogas. The shiny application was used to determine various pa-rameters in biogas simulation and the screenshots of the application are shown infigure 2 obtained from https://biotransformers.shinyapps.io/oba1/ .
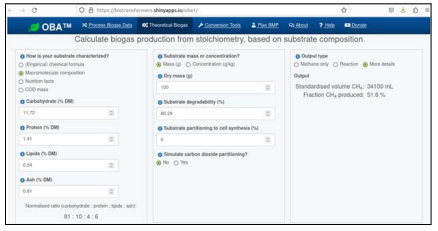
Figure 2: A screenshot of online Biogas App
The obtained results for crude fat, proteins, fiber and carbohydrates are shown in table 1. High crude fat content was recorded in avocado at 9.03% while protein was lowest in tomato at 0.57%. Low crude protein content in fruits had earlier been observed by at 0.09 - 3.54% range [30].
| Sample | % Ash Content |
% Protein |
% Fat | % Carbohydrates |
|---|---|---|---|---|
| Kales | 1.94 | 2.27 | 0.34 | 4.03 |
| Cabbage | 0.49 | 0.83 | 0.05 | 3.22 |
| Pumkin Leaves | 2.06 | 2.27 | 0.18 | 3.77 |
| Cucumis ficifolia | 2.34 | 3.49 | 0.33 | 5.74 |
| Pigweed | 2.86 | 2.61 | 0.21 | 3.62 |
| Erucastrum arabicum | 1.99 | 2.82 | 0.19 | 3.95 |
| Coriander | 1.91 | 2.6 | 0.09 | 2.16 |
| A. nightshade | 1.97 | 2.68 | 0.26 | 4.12 |
| Spinach | 1.73 | 1.53 | 0.17 | 2.38 |
| Comfrey | 3.46 | 3.24 | 0.29 | 5.9 |
| Tomato | 0.46 | 0.57 | 0.12 | 2.93 |
| Potato | 0.81 | 1.41 | 0.54 | 11.72 |
| Sweet Potato | 1.06 | 1.67 | 1.54 | 32.17 |
| Pawpaw | 0.5 | 0.68 | 0.34 | 7.95 |
| Banana | 1.67 | 3.05 | 0.5 | 19.24 |
| Avocado | 0.84 | 1.32 | 9.03 | 3.37 |
| Courgette | 0.72 | 1.06 | 0.25 | 1.99 |
| Cucumber | 0.46 | 0.52 | 0.21 | 2.17 |
| Mango | 0.44 | 0.87 | 0.68 | 9.91 |
| Water Melon | 0.74 | 0.9 | 0.33 | 4.42 |
| Waste Mixtur | 1.4225 | 1.8195 | 0.783 | 6.738 |
Standard deviation ranges: 0.03-0.96 protein, 0.01-0.69 fat, 0.24 -0.85 fiber and 0.33 -1.02 carbohydrates.
The proximate composition of the substrates showed that carbohydrates levels was higher in all the feedstocks compared to proteins and fats. This is because sugars form the fundamental blocks in most tissues which further translates to higher energy/100g of each waste.
The nutrient content analysis showed that the food waste contained well balanced nutrients for anaerobic microorganisms. The results of this study indicate that the food waste is a highly desirable substrate for anaerobic digesters with regards to its high biodegradability and methane yield which had earlier been reported by [31].
The results of biodegradability of the market waste sample are shown in table 2. Digestibility shows the ability of a substrate to degrade.
| Substrate | BD exp | BD LB | BD ele |
|---|---|---|---|
| Kales | 86.24±2.34 | 77.40±1.09 | 83.91±2.11 |
| Cabbage | 83.19±l.00 | 73.90±1.20 | 80.50±1.53 |
| Pumkin Leaves | 84.64±2.19 | 75.72±3.01 | 79.60±l.00 |
| Cucumis ficifolia | 77.36±3.99 | 71.81±3.00 | 81.98±1.54 |
| Pigweed | 85.88±0.99 | 68.36±1.26 | 81.51±1.64 |
| Erucastrum arabicum |
76.85±5.87 | 73.07±1.88 | 73.07±1.88 |
| Coriander | 85.43±0.89 | 69.28±2.89 | 77.83±1.93 |
| A.nightshade | 80.77±2.33 | 74.04±1.66 | 76.57±1.07 |
| Spinach | 80.00±l.50 | 74.88±2.06 | 77.80±0.87 |
| Comfrey | 86.96±7.00 | 71.80±1.96 | 78.64±1.90 |
| Tomato | 72.92±3.33 | 72.92±2.00 | 79.36±l.98 |
| Potato | 84.10±2.12 | 76.56±1.19 | 80.22±1.22 |
| Sweet Potato | 84.82±7.88 | 70.43±2.36 | 77.75±0.68 |
| Pawpaw | 84.73±5.63 | 76.56±1.88 | 77.04±0.89 |
| Banana | 86.27±5.73 | 74.04±2.05 | 75.74±1.45 |
| Avocado | 84.08±0.82 | 76.56±2.76 | 77.36±0.95 |
| Courgette | 77.16± 1.26 | 68.16±1.33 | 74.83±1.55 |
| Cucumber | 78.26±3.56 | 69.28±1.25 | 76.16±l.67 |
| 79.07±1.88 | 77.12±2.89 | 81.95±0.99 | Mango |
| Water Melon | 76.61±.32 | 71.82±l.22 | 74.65±1.00 |
| Mix | 93.48±l.11 | 77.76±1.26 | 83.51±1.78 |
BD ele describes the degree to which biomass is degraded and categorized into biodegradable and non-biodegradable. Due to undigested VS and chemical oxygen demand, the BDexp was determined using equation 13. Similarly, in comparison with BD exp and BDeleas described in table 2, lignin-based (BDLB) has shown the least degradability of the substrate. The BD order depended heavily on the fiber content of the lignin matter of the substrate. Lignin’s existence determines the degree of BD and biogas yields. This further implies that other organic content relates directly to biogas generation. It is observed that BDele was lower than BDexp and BDLB. The high levels of BDexp means that some volatile matter is utilized for microbial growth and metabolism. The relationship between BD and lignin has shown that 1 percent of lignin reduces biomass degradability by about 3-fold. Slight differences in BD and BMP results are also observed, which may vary due to different plant conditions appropriate for AD i.e., temperature, flask gas-space size, inoculum, inoculum substrate ratio.
The characterization of solid wastes is a necessary step before they can be used in anaerobic digestion. The quantities of different compounds (carbohydrates, proteins, lipids, and fibers) and anaerobic biodegradability (capacity to produce methane) are important information required to characterize waste [32]. Theoretical biogas yields largely depend on lipids, carbohydrates, and proteins levels [33]. The main proximate properties involve analysis of moisture, carbohydrates, protein, and fat content.
The carbohydrates levels were reported to be the highest among the proximate properties investigated in this study. The highest amount was reported in sweet potato at 32.17 % and lowest in courgette at 1.99 %. The carbohydrate or the complex sugars are broken down into monosaccharides e.g., lactose into galactose and glucose as shown in equation 10.0.

It is evident from table 3 that biogas generation is highly dependent on the carbohydrates level in the sample. It was observed that at high carbohydrate levels, high biogas was generated [34]. The fat levels amongst the substrates were higher in avocado at 9.03 % and lowest in kales and cabbages at 0.09 and 0.05 % respectively.
| Sample | % Digestibility |
Standardized methane (L) |
Fraction CH4 produced (%) |
|---|---|---|---|
| Kales | 82.52 | 32.24 | 52.41 |
| Cabbage | 79.2 | 31.53 | 50.90 |
| Pumkin Leaves | 79.99 | 28.52 | 51.82 |
| Cucumis ficifolia |
77.05 | 29.64 | 52.02 |
| Pigweed | 78.58 | 26.21 | 52.01 |
| Erucastrum arabicum |
74.01 | 27.60 | 51.81 |
| Coriander | 77.51 | 27.20 | 51.83 |
| A. nightshade | 77.13 | 29.01 | 52.12 |
| Spinach | 77.56 | 26.32 | 52.21 |
| Comfrey | 79.13 | 27.41 | 51.84 |
| Tomato | 78.16 | 31.41 | 51.60 |
| Potato | 80.29 | 34.12 | 51.61 |
| Sweet Potato | 77.67 | 33.63 | 51.63 |
| Pawpaw | 79.44 | 33.54 | 51.53 |
| Banana | 78.68 | 32.51 | 51.11 |
| Avocado | 79.33 | 61.12 | 65.52 |
| Courgette | 73.38 | 30.01 | 53.12 |
| Cucumber | 74.57 | 30.81 | 52.82 |
| Mango | 77.91 | 34.42 | 52.23 |
| Water Melon | 74.36 | 30.82 | 52.31 |
| Waste Mixture | 84.92 | 35.91 | 53.12 |
It was also noted that the fat content influenced biogas generation in waste to biogas conversion. High-fat levels translated to high biogas production. The overall biogas produced by the avocado substrate was in 600-2600 mL range for the seven days’ retention time. Fat is converted to fatty acids in the hydrolysis step as described below in the reaction proposed by [35].
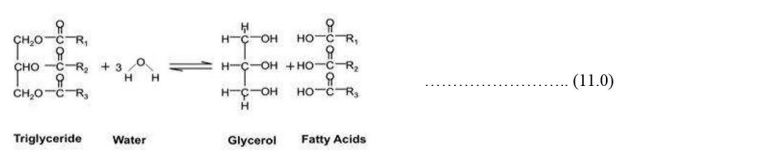
Among the proximate matter, lipids contribute largely to biogas formation though with longer HRT because of slow bio-degradability. Proteins and carbohydrates have fast digestion rate though the yield is low [33].
The protein levels were lowest in tomato wastes at 0.58 and highest in Cucumis ficifolia wastes at 3.49 %. The observed trend is that the higher the protein levels, the lower the biogas production.
The protein content influences the levels of ammonia and hydrogen sulfide in the digester. This translates to some inhibition of microbial activities, consequently influencing biogas productions. The resulting equation is shown in equation 12.0.

studied the effects of carbohydrates, protein and fat on biogas generation using vegetable waste, oil cake and whey [36]. They observed that methane production was dependent on these proximate proprieties. This was also reported by [36,37]. With a fixed slurry concentration, methane levels decreased with an increase in carbohydrates concentration because at high levels of carbohydrates, acidogenic bacteria growth is favored producing volatile fatty acids like butyric and valeric which inhibit methanogens growth and therefore, low methane generation. Besides, high protein content leads to low methane formation due to the formation of ammonia at the acetogenesis step [36]. On the other hand, fat content favors methane production due to the availability of long fatty acids being converted to methane [38,39]. reported biogas yield of 790, 1250 and 700 L/Kg of organic matter and methane levels of 50, 68 and 71 % for carbohydrates, fats and proteins respectively [40-43].
The application of anaerobic digestion technology in conversion of market wastes to renewable energy and bio-slurry is key in managing organic landfill wastes. The proximate composition of the substrates showed that carbohydrates levels was higher in all the feedstocks compared to proteins and fats. This is because of sugars form the fundamental blocks in most tissues which further translates to higher energy/100g of each waste.
The authors wish to express their sincere gratitude to National Research Fund (NRF), grants no. 501-000-053 for funding this research work.
